Data access
Data from DanMAX is stored centrally at MAX IV and is available for further processing using MAX IV resources – or for download using either the Globus connect service or sftp. Detailed information about downloading the data can be found at the following link.
Data handling guides
Format
All scans at DanMAX results in a number of files written in the HDF5 format (*.h5). The files are written to be NeXuS and/or DXchange compatible.
Destinations
The raw files are stored in:
/data/visitors/danmax/#Proposal/#Visit/raw/while processed files are stored in:
/data/visitors/danmax/#Proposal/#Visit/process/Ipython notebookes for processing data will be placed in:
/data/visitors/danmax/#Proposal/#Visit/scripts/Separate experiments/samples may be stored in their own folder. The raw folder is write protected to protect the raw data. The process, scripts, and other folders are not protected, so it is possible to edit/overwrite files in these
Archiving
Each scan will be saved with the name scan-####.h5 where #### is a sequential number. It is important to keep track of this number and the corresponding sample/experiment, e.g. by using the Elogy logbook at elogy.maxiv.lu.se.
The scan-####.h5 file is a master file that contains the command issued to perform the scan, the scanned parameters (e.g. motor positions), experimental channels (e.g. beam intensity, position, temperature) a snapshot of the instrument at the beginning of the scan and links to the detector data files.
The data from the detectors are located in separate files, e.g. scan-####_pilatus.h5, scan-####_falconx.h5, and scan-####_orca.h5. The link from the master file to the detector data is relative, and thus all files must be located in the same folder.
The tomography data is also saved in the DXchange format for straightforward processing using available scripts incorporating, e.g., TomoPy. This file is located in the raw folder with the name scan-####_dxchange.h5. This file still links to the detector data located in separate files.
Additional information
To visualize the data and the file structure, we recommend using silx view, which is available on the control computers. The Silx installation files can be found on silx.org
As the PXRD area detector data is compressed using a bit-shuffle algorithm, it is necessary to install hdf5plugin, which can be found on pypi.org
A description of the experimental snapshot parameters is given in the next tab.
Azimuthal integration pipeline
An automated azimuthal integration pipeline is available based on the MATFRAIA algorithm developed by the Birkedal group at Aarhus University (Publication: https://scripts.iucr.org/cgi-bin/paper?S1600577522008232, Python source code). To prepare the pipeline, a .poni file and a detector mask (.npy) are needed.
pyFAI-calib2 is installed on the control computers and is commonly used to generate these files. More information about the pipeline can be found on the beamline wiki, under Pilatus 2M CdTe
The integrated data will be available as saved in the following folder, mirroring the folder name and scan number in the /raw/ folder.
/data/visitors/danmax/#Proposal/#Visit/process/azint/From the fall of 2025, the azimuthally integrated data are saved in the NXaxint definition (Normal 1D data: NXazint1d, Azimuthally binned data: NXazint2d). We are working with different software developers to implement support for these files directly in some of the most popular PXRD refinement programs – stay tuned for updates!
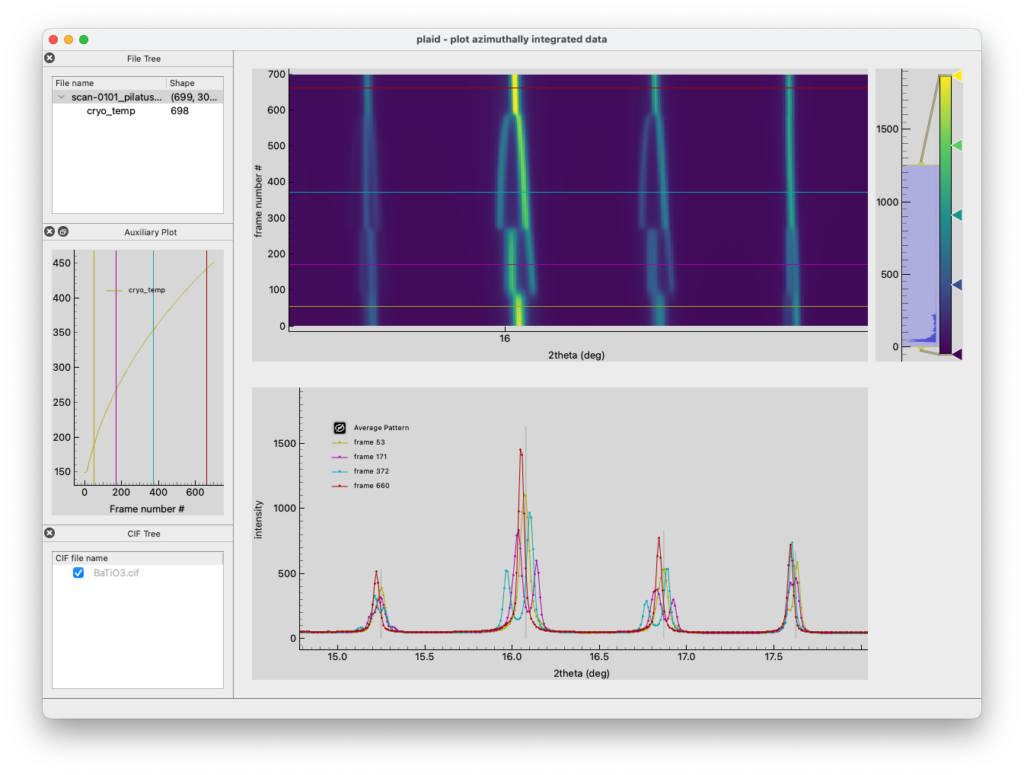
To visualize the files (and potentially export the data as ASCII XY files), we have developed the program ‘plaid’ (‘plaid looks at integrated data’ – formerly ‘plot azimuthally integrated data’). More information can be found here https://pypi.org/project/plaid-xrd/. Install using: pip install plaid-xrd
Jupyter
It is also possible to process the data in Jupyter notebooks using the MAX IV Jupytherhub, where you need to log in using your DUO credentials. To have access to the regular environment with HDF5, silx, and pyFAI tools, you should check the “Beamline-specific analysis” radio button and select “DanMAX […]” from the drop-down menu.
Reconstruction pipeline
An automatic tomographic reconstruction pipeline is available and described in this paper: Christensen et al., J. Synchrotron Radiat. (2026).
To prepare the pipeline, a .tori (Tomographic Reconstruction Information) file is needed. This file is prepared in the ‘Reconstructor’ GUI available at the beamline.
The reconstructed volumes will be available as saved in the following folder, mirroring the folder name and scan number in the /raw/ folder.
/data/visitors/danmax/#Proposal/#Visit/process/recon_pipeline/Jupyter
It is also possible to process the data in Jupyter notebooks, using either the normal MAX IV Jupyterhub (~8 cores, 100 GB of memory) or the high-performance HPC Jupyterhub (Up to 32 cores, 384 GB of memory and possibility for Nvidia V100 GPU). You will need a VPN connection and DUO credentials to log in to either system.
Select the “Tomography” -> “tomorec” kernel for access to the correct packages for tomographic reconstruction and analysis.
Image Analysis Support from the Qim Center
Following the acquisition of your data, you can leverage the tools and expertise offered by the Center for Quantification of Imaging Data from MAX IV (Qim Center) to support your analysis process.
The Qim Center provides the following resources:
- Advanced methodologies specifically designed for volumetric data.
- A Python library (qim3d) that delivers comprehensive functionality for volumetric image analysis.
- Access to the Qim Platform, a web-based interface facilitating the utilization of HPC resources at DTU.
For examples of the methodologies developed by the Qim Center, please visit Qim Software Tools.
Additionally, the Qim Center’s team of specialists is available for consultation to optimize your data analysis approach.
You can contact the QIM team by emailing info@qim.dk or contacting one of the specialists directly.
The experimental snapshot is a recording of the essential instrument parameters at the beginning of the scan. The snap shot is contained in the master file for each scan, and is located under
/entry/instrument/start_positioners/.| Name | Description |
| IVU_R3_304_GAP | Is the undulator gap |
| hdcm_energy | Energy of the Si111 hDCM based on encoder position |
| mlm_energy | Energy of the hMLM based on encoder position |
| *_slit_hgap/hoff/vgap/voff | Is the horizontal/vertical openings and offsets of the slits |
| bcu_* | Is referring to positions of the equipment on the Beam Conditioning Unit |
| tom_* im_* nf_* | Is referring to positions of the axis of the Tomograph |
| ben_* sg_* | Is referring to positions of the axis of the optical bench and the small gantry |
| pd_* | Is referring to the positions of the equipment on the PXRD2D instrument |
| lg_* | Is referring to the position of the Large Gantry |
| bcu_xbpm_x/y | Is the X and Y position of the X-ray beam position monitor on the BCU |
| nf_z | The tomography sample to microscope distance. Caution: May not reflect the actual distance if a calibration has not been performed after changing the configuration. |
| pd_pinhole_x/y | Is the X and Y position of the PXRD2D pinhole |
| pd_hex_x/y/z/rx/ry/rz | Is the position and angle of the PXRD2D hexapod |
| pd_huber | Is the position of the optional rotation table at the PXRD2D instrument |
| pd_spinner_moving | Was the PXRD2D sample spinner spinning at the beginning of the scan? |
| pd_bst_x/y/z | Is the location of the PXRD2D beamstop |
| lg_z | PXRD2D sample to detector distance (Pilatus 2M on the large gantry) of it is not tilted. Caution: May not reflect the actual distance if a calibration has not been performed after changing the configuration. |
| Pilatus energy | Is the energy setting on the Pilatus responsible for setting the flatfield of the detector |
| Pilatus threshold | Is the threshold energy of the Pilatus responsible for energy discrimination |
| pd_xrf_z | PXRD2D sample to XRF detector distance. Caution: May not reflect the actual distance if a calibration has not been performed after changing the configuration. |
Some names are self-explanatory and not listed here.